The Sequence 4/17-4/23
Insights Into the Deciphering Developmental Disorders Study, Treatment for Adrenoleukodystrophy, Polygenic Risk Scores in Predicting Coronary Artery Disease, Insights Into Macular Degeneration
New insights into the Deciphering Developmental Disorders study
Researchers at Exeter and the Wellcome Sanger Institute among others in the UK and Ireland joined together in 2011 to participate in the Deciphering Developmental Disorders (DDD) study. The goal was to find genetic diagnoses for thousands of patients with difficult-to-diagnose developmental disorders.
How did they do that?
The monumental DDD study collected clinical information from and performed exome sequencing and microarray analyses on more than 13,500 families in which patients had severe, difficult-to-diagnose developmental disorders. They then developed a pipeline to find genetic variants that were likely the cause of these disorders, and reported those variants back to clinical teams.
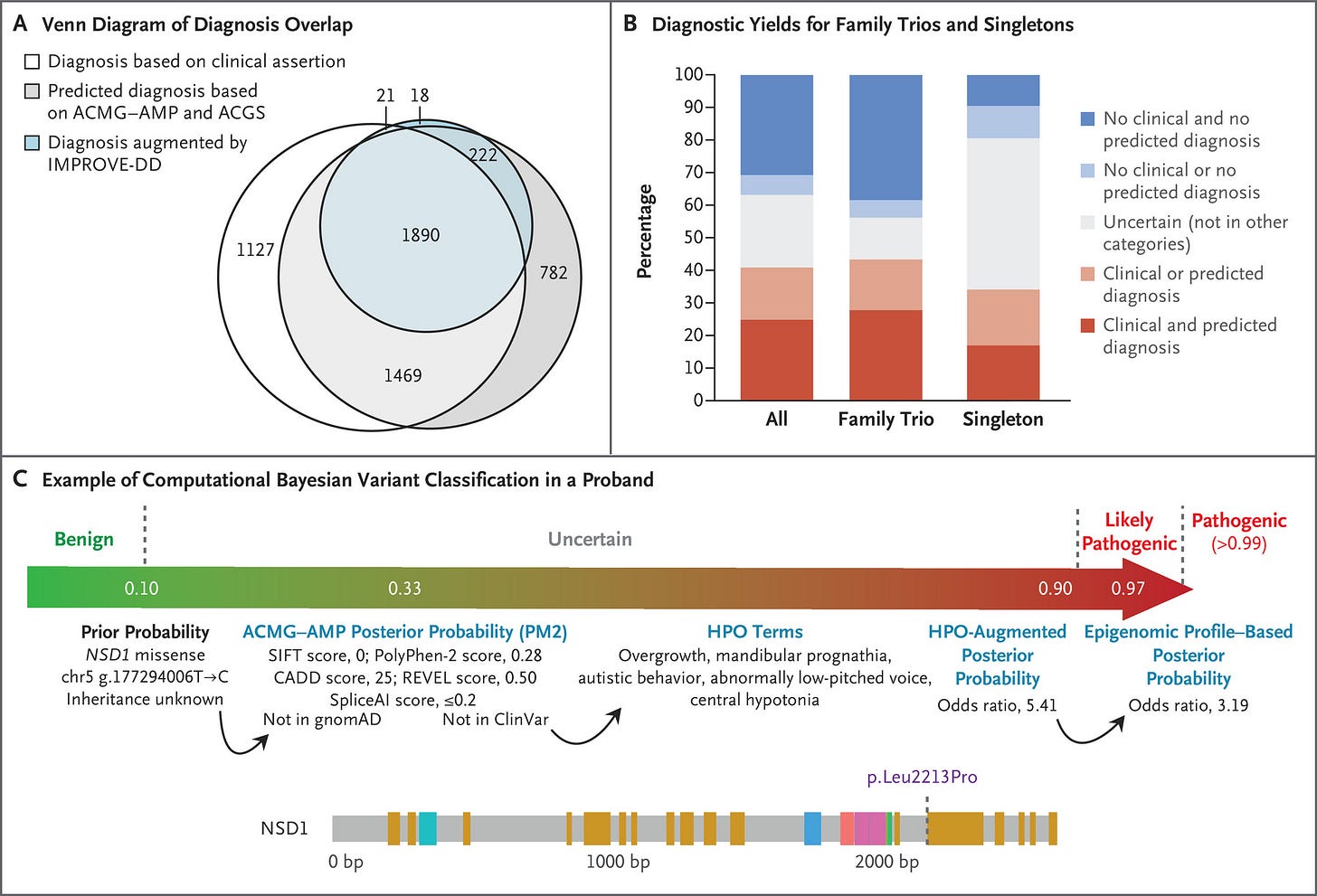
Diagnoses in the DDD Study, figure by Wright et al. 2023
Awesome. What were the results?
Of the 13,449 participants whose data were analyzed, a diagnosis was made in approximately 41% of patients. Of those diagnoses, 76% were pathogenic de novo, meaning they were causative genetic variants that were not inherited from the patients’ parents.
When looking at what factors may affect the probability of getting a genetic diagnosis, they found that inclusion of parents in testing had the largest effect on increasing the probability of diagnosis. A history of premature birth (i.e., 22 to 27 weeks’ gestation), in utero exposure to antiepileptic medications, mothers with diabetes, and African ancestry decreased the probability of a genetic diagnosis (see more on why ancestry makes a difference in this previous post).
Additionally, the analysis pipeline that was developed to find causative genetic variants, shown in panel C of the figure above, successfully incorporated both genetic and clinical data in order to best classify genetic variants identified.
That’s a lot of great information.
It is! This cohort is sort of a big deal. I can’t tell you how many times I’ve used DECIPHER, the database where results from DDD are housed, in my quest for gene discovery as a genetic counselor working in research. The DDD study is one of the largest translational genomics studies to date, and particularly when we’re talking about studies of patients with developmental disorders. All in all, this study points to a large diagnostic yield from genetic testing and detailed clinical phenotyping, and supports the potential for diagnoses through genetic testing for patients with developmental disorders. The more we utilize this cohort in research, the more we will learn in the years to come.
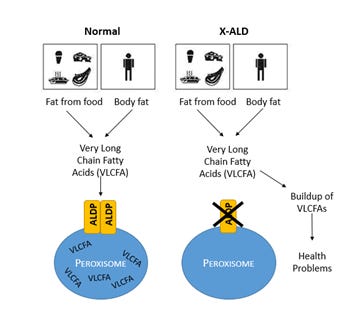
Disease mechanism of X-linked adrenoleukodystrophy, Image credit: newbornscreening.info
Gene therapy as a treatment for X-linked adrenoleukodystrophy
X-linked adrenoleukodystrophy (ALD) is a disease in which the most severe form causes a sudden onset of learning and behavioral problems, and often total disability and death within only a few years. This is caused by the buildup of very long-chain fatty acids (VLCFAs) in the body as they fail to enter an organelle called the peroxisome. This buildup of VLCFAs results in damage to cells in the brain.
The good news: There is treatment for X-linked ALD. Treatment is usually hematopoietic stem cell transplantation (HSCT), which involves some serious risks including those related to surgery preparation, the transplantation itself, and then infections and failure post-transplant.
How does the current treatment work?
HSCT replaces cells in the body, allowing cells to enter the brain in replace of cells that have been damaged by the build-up of VLCFAs.
What is the new treatment?
Gene therapy! Lauer et al. reported that HSCT using cells containing gene-editing technology by way of a viral vector (read more on that here) caused a normalization of white matter. Important, as normal white matter patterns are usually disrupted by damaged cells in the brain of patients with X-linked ALD.
Once infused into the bone marrow, the gene-editing technology allows for the editing of the ABCD1 gene, the gene harboring mutations in patients with X-linked ALD, allowing for functional adrenoleukodystrophy protein and therefore efficient breakdown of VLCFAs.
What’s the takeaway?
Not only were gene-edited cells derived from HSCT able to insert themselves into the important parts of the brain needed to increase function in patients with X-linked ALD, evidence suggested these changes would continue to increase function long-term. This marriage of HSCT + gene therapy may be the ticket to an elevated and more effective treatment for those with X-linked ALD.
The utility of polygenic risk scores in predicting coronary artery disease
A study by Örd et al. examined data on mice with and without coronary artery disease to evaluate the accuracy of polygenic risk scores (PRS) that take into account different cell states in predicting cardiovascular disease.
What do you mean different cell states?
A cell state refers to the various ways a cell can respond to a certain condition. For example, in response to inflammation, cells may express differently, leading to the circulation of different types of immune cells. In this article, the research team wanted to know whether different levels of cell states may signify different likelihoods of coronary artery disease.
What is a polygenic risk score?
A polygenic risk score (PRS) is a number, or a ‘score’ that estimates an individual’s risk for a certain condition. They are used in conditions that are caused by changes in many genes, often coupled with environmental factors.
How are they determined?
Let’s use coronary artery disease as an example, because that is the health outcome studied in this article. Scientists created PRSs for coronary artery disease by comparing the DNA of patients with and without coronary artery disease to determine a ‘collection of genes’ that have more rare variation in the individuals with coronary artery disease that are not there in the individuals without coronary artery disease. Then, they can say if you have these ‘X’ number of genes, your PRS for coronary artery disease is increased.
What did they find?
The group did in fact find that genetically predicted risk for 12 different disease-associated cell states (i.e. based on a high PRS) was associated with cardiovascular risk.
What's the takeaway?
I talked about PRS in association with cardiovascular disease risk here. This time, however, we’re getting more granular. We’re not only talking about a score to predict a clinical symptom, we’re talking about a score to predict what’s going on in the body at a cellular level. It would be pretty amazing to be able to use a PRS (which is calculated from only a blood sample, by the way) to identify risks for cardiovascular disease; especially in patients without any other clinical indicators. Understanding our risks for common conditions such as cardiovascular disease can guide healthcare management.
Insights into macular degeneration
I’ve had conversations with more than one field leader who view multi-omics as the future of genetics. In this study, Orozco et al. used multi-omics to understand the underlying cause of age-related macular degeneration (AMD).
Tell me more.
Age-related macular degeneration (AMD) is the most common cause of vision loss among older adults, affecting 200 million people worldwide. It is caused by damage to the central part of the retina. Currently, there are a couple of well-known genetic variants that cause an increased susceptibility to AMD- those are genetic variants in the ARMS2 and CFH genes.
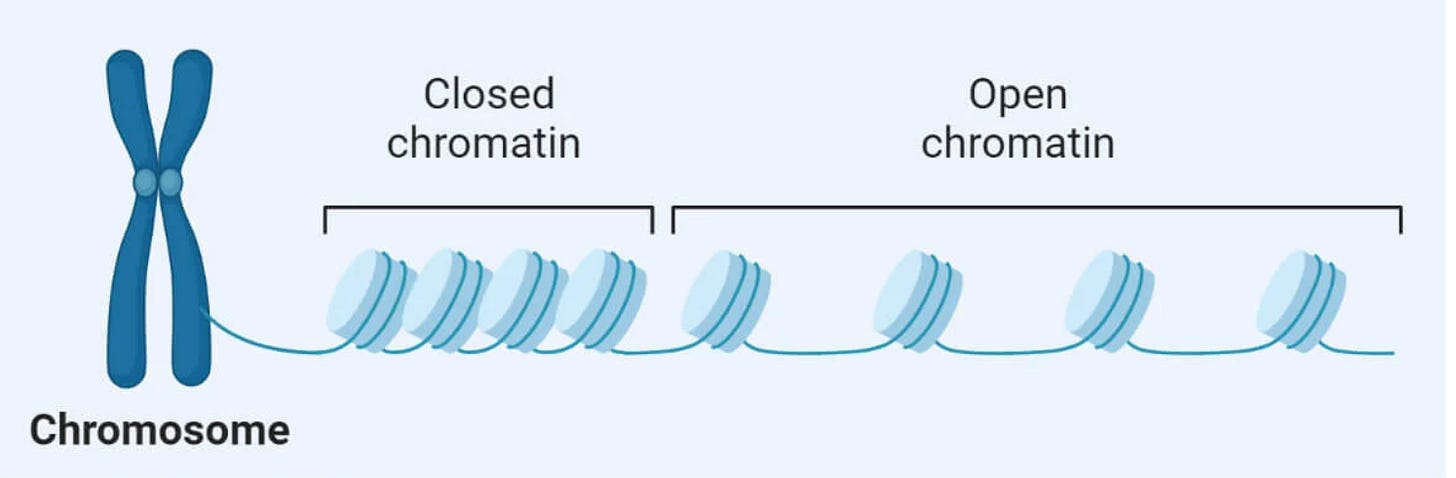
Chromatin, image credit: thebiologynotes.com
Got it. What did the genetic analysis of the ‘omics find?
The team analyzed different regulatory elements in cells from both the outermost layer of the retina as well as the choroid, or the layer of connective tissue under the retina, from patients with and without AMD. In the group with AMD, they identified changes in genes associated with chromatin accessibility, literally the accessibility of chromatin, which is made up of DNA and proteins (it’s the stuff that packages up chromosomes). The more accessible chromatin is, the more easily it is able to interact with other proteins, like transcription factors, to control gene expression. All this to say, genes associated with chromatin accessibility and gene expression, including HTRA1 and C6orf223, may play a large role in the development of AMD.
Additionally, they identified a couple of important genes active in the WNT pathway, FRZB and TLE2, indicating the importance of this particular cell signaling pathway in AMD.
What’s the takeaway?
This study marks a major step toward understanding the mechanism behind AMD, including what genes and complex pathways are involved in its onset. Understanding the genes and mechanisms involved in certain conditions opens up options for therapy in the future.
In other AMD news, this study identified causative genetic variants in 5 genes associated with immune response pathways.