The Sequence 5/29-6/4
Engaging a Community With a History of Huntington’s Disease, A Cheaper Way to Sequence DNA, New Animal Species Discovered Deep in the Pacific Ocean, The Anti-pain Gene
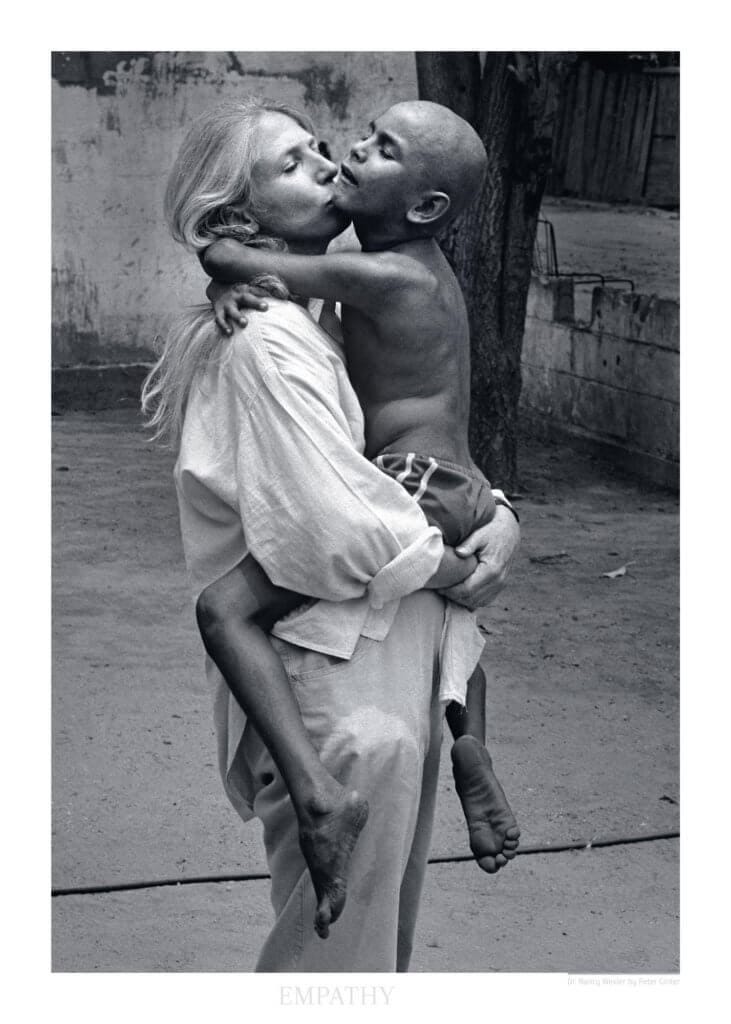
Nancy Wexler with Eduardito, age 15, Zulia, Venezuela; Image credit: Peter Ginter, originally published in the NYT
Engaging a community with a history of Huntington’s disease
Nancy Wexler, the ‘mother of Huntington’s’ as I've heard her described, traveled to Lake Maracaibo, Venezuela, in 1975 on a mission to find the cause of Huntington’s disease. In her time there, she collected over 4,000 blood samples and documented 18,000 different individuals to work out a common pedigree. Really, though, she did more than collect some data. She became a part of the community of people there. She gave them hope for a treatment, and hope for all of the good that this research could do. And she was right; the data collected from Lake Maracaibo led to the discovery of the location of the gene associated with Huntington’s Disease by 1983.
Importantly, Lake Maracaibo was not the only place where research was being conducted on large families with Huntington’s disease. Similar studies were taking place in neighboring Colombia under the direction of Dr. Jorge Daza, a local neurologist based in Barranquilla.
But what happened to the families that were studied by these pioneers, and who inspired generations of scientists to come? Here, I discuss an article by Jennie Erin Smith of the New York Times updating us on the people living in Colombia who participated in research by Dr. Daza.
How are they?
A bit neglected, if we’re being honest. It seems like once Dr. Daza passed away, everything stopped abruptly. No one took over his work, and no one continued to treat and care for these families. Once symptoms start for these patients, they do not know where to go to get treatment or what the treatment is. In some cases, even the physicians who care for these families do not know what Huntington’s disease is. One patient quoted in the article described having to write out the name of the disease for her doctor.
Is anyone doing anything about it?
They’re trying. A group of researchers at the Universidad Simón Bolívar, in Barranquilla, have begun the process of re-finding these families in order to continue research and provide treatment. But that’s just it: they had to restart communication by even finding them in the first place.
At this point, the team has successfully educated local health professionals, and they’re optimistic about the prospects of enrolling the families into Enroll-HD, a global platform to study people with Huntington’s disease and facilitate clinical trials.
Awesome. What’s the takeaway?
As someone who was personally inspired by Nancy Wexler’s work, I feel it’s important for the new generations of scientists to mirror admirable work like Wexler’s and Daza’s- the societal investment in the places that are essential to research- and improve on it even more. It’s of utmost importance to pay attention to the inclusion and engagement needs of understudied populations, and to make that valuable relationship a long-term one.
Full article; Resource funding current Huntington’s Disease research: The Hereditary Disease Foundation (HDF)
Another article revealing perspectives from an individual carrying a pathogenic variant that causes Huntington’s Disease.
A cheaper, more efficient way to sequence DNA
As DNA sequencing technology gets better and processes are more streamlined, the cost is rapidly decreasing. We have certainly come a long way from the $2.7 billion that was required to first sequence all of the genes in the human genome for The Human Genome Project. However, DNA sequencing is by no means ‘cheap’.
That’s where a new sequencing technology called avidity sequencing comes in. This week, Sinan Arslan and colleagues presented data on the incredible accuracy of this new sequencing technology.
How does avidity sequencing work?
Let’s begin by talking about how other sequencing technologies work. The most commonly used method, called ‘sequencing by synthesis’, works like this:
DNA bases (A, C, G, T), each of which are labeled with a different color, are flowed across a solid surface, to which sample DNA is attached. As the bases in the flowing reagent come across bases in the sample that are complementary, they attach. A picture is then taken that reads the colors that have attached to the solid surface, and voila: we have a sequence of letters based on translating the colors. In order to read a whole sequence, this process is repeated many times.
Now, for avidity sequencing:
In avidity sequencing, the flowing reagent contains avidite substrates. The avidite substrate has the ability to bind multiple bases of DNA at one time. This reduces the concentration of flowing reagent needed, overcoming sequencing cost barriers.
Makes sense.
It does. And according to the study by Arslan et al., avidity sequencing was pretty accurate, with 96.2% and 85.4% of base calls having an average of one error per 1,000 and 10,000 base pairs, respectively.
What’s the takeaway?
Sequencing by synthesis has been the gold standard in large scale sequencing for quite a while, and certainly as long as I've been in the field. Historically, other types of technology that have come along, like long-read sequencing for example, have been considered very costly and full of errors. But it seems like that’s changing. I look forward to seeing the opportunities avidity sequencing brings in light of its fast and accurate generation of data.
Interested in more ways genetic testing is becoming more cost effective? This article discussed combining three common genetic tests for young people into one.
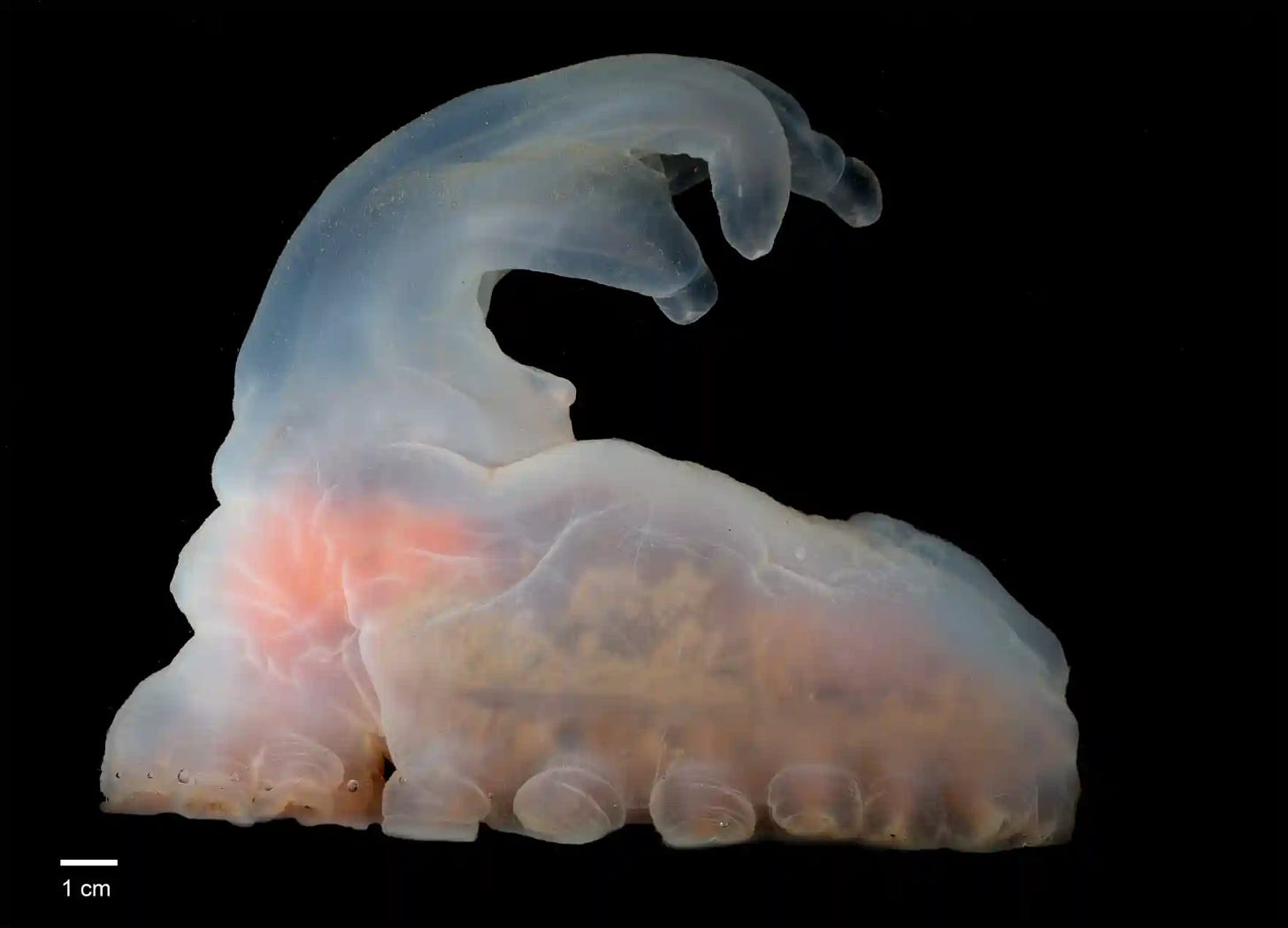
The Amperima, image credit: Explorersweb
New animal species discovered deep in the Pacific ocean
More than 5,000 new species have been discovered in the Clarion-Clipperton Zone (CCZ), an area of the Pacific ocean floor. Important, because the mineral-rich area is contracted for deep-sea mining, which could mean extinction of these new species.
In order to encourage effective management of environmental impact from potential deep-sea mining activities, Rabone et al. published the first checklist taking inventory of the species identified in the CCZ.
Why is a checklist important?
Firstly, of the 5,578 new species recorded, an estimated 88%–92% of them have not even been described before. This means they have not been categorized into phyla, which are organized by shared structural features and/or shared ancestry. This is where DNA analysis comes in.
By being able to sort these species into the appropriate phyla based on ancestry and DNA analysis, we can answer questions like how impacted some of these organisms that live in the mineral-rich sediment may be by mining.
What’s the takeaway?
Given the mining operations planned in the CCZ, the CCZ checklist is an important step toward considering the biodiversity in the area. By understanding what organisms are living in the sediment, we can take proper steps toward preventing disruption of the environment, which could include harm of the current organisms living there and even extinction.
Interested in more articles discussing the impact of genetic research on understanding the origins and classifications of animals? These three studies discuss the evolution of marine sponges, butterflies, and wooly mammoths.
The anti-pain gene
Hajar Mikaeili and colleagues studied the DNA of a patient with pain-insensitivity to understand if there is a genetic difference that may be causing the insensitivity to pain. They identified a deletion in the FAAH-OUT gene that is likely the cause.
Interesting. What does the FAAH-OUT gene do?
The FAAH-OUT gene is part of a pathway that is involved in pain modulation, anxiety and stress responses, memory, and wound healing. The idea is that a deletion in the FAAH-OUT gene would disrupt proteins in that pathway, meaning those controls may be atypical. Interestingly, the patient that was identified with pain-insensitivity also had reduced anxiety and fast wound healing.
The team then performed functional studies using patient cells to see what affect the deletion has on a molecular level. Those studies showed that the deletion in the FAAH-OUT gene reduced expression of another important gene in that pathway called FAAH.
What’s the takeaway?
This new understanding of the role of the FAAH-OUT gene could mean a way to target treatment of pain, anxiety, depression, and other neurological disorders on a molecular level. The possibility of permanent treatment could be life-changing for the population of people struggling with these conditions.
In other therapy news, an update on my article from last month on the approval of Sarepta Therapeutics’ gene transfer therapy for Duchenne muscular dystrophy (DMD): The FDA needs more time.